
Center for Optimized Oligo Escape and Control of Disease (COE)
COE is a challenge center funded by the Novo Nordisk Foundation for a 6-year period.
The scope of the Novo Nordisk Foundation - Center for Oligo Escape and Control of Disease is to provide the required mechanistic understanding to optimize subcellular delivery of RNA and other oligonucleotides entrapped in endosomes, develop methods for their efficient cellular delivery, and demonstrate applicability by targeting skeletal and cardiac muscle cells.
The vision of COE is to resolve a main bottleneck that prevents the widespread implementation of oligonucleotide pharmaceutics, and their limited (~<5%) endosomal escape.
The Center is located at the University of Copenhagen with two satellite laboratories, one at Harvard Medical School & Boston Children’s Hospital, the other at Instituto de Medicina Molecular, University of Lisbon.
COE combines the interdisciplinary expertise of leading experts in oligonucleotide synthesis (Jensen), with studies at the fundamental limit of individual biomolecules (Hatzakis), high spatiotemporal precision live cell imaging (Kirchhausen), machine learning based computational image analysis (Hatzakis, Kirchhausen), and ex-vivo splicing modulation to correct skeletal and cardiac muscle pathologies (Fonseca).
The provided knowledge will iteratively improve the oligonucleotide conjugate modifications and delivery methods required to overcome the limited endosomal escape of oligonucleotide pharmaceutics initially evaluated to systems relevant for cardiometabolic diseases.
Besides creating a bridge between the leading laboratories and institutions, the complementary interdisciplinary expertise and scientific impact of our team will create a premier multidisciplinary research environment for innovative studies on ways to improve the cytosolic delivery of functionally active RNA and other oligonucleotides targeting a spectrum of diseases.
Hatzakis lab
Hatzakis is leading the development and implementation of single-particle in vitro assays and in vivo cell studies to study the interplay of spatiotemporal localisation and function of biomolecular entities (protein and oligonucleotide) and their role in cellular responses. His group is actively implementing machine-learning pipelines for automatic, rapid, and precise extraction of quantitative information from fluorescence microscopy data.
Read more about the Hatzakis Lab on on the UCPH website.
Selected papers:
-
Nature Chemistry 2022 14(5), 558-656
Mette Galsgaard Malle, Philipp M. G. Löffler, Søren S.-R. Bohr, Magnus Berg Sletfjerding, Alexander Risgaard, Simon Bo Jensen, Min Zhang, Per Hedegaard, Stefan Vogel, Nikos S. Hatzakis*
“Single particle combinatorial multiplexed liposome fusion mediated by DNA” -
Nature Commun 2021 12, 2260
Simon Bo Jensen, Sara Thodberg, Shaheena Parween, Matias E. Moses, Cecilie C. Hansen, Johannes Thomsen, Magnus B. Sletfjerding, Camilla Knudsen, Rita Del Giudice, Philip M. Lund, Patricia R. Castaño5,6, Yanet G. Bustamante1, Maria Natalia Rojas Velazquez, Flemming Steen Jørgensen, Amit V. Pandey, Tomas Laursen, Birger Lindberg Møller, Nikos S. Hatzakis*
“Across kingdom biased CYP-mediated metabolism via small-molecule ligands docking on P450 oxidoreductase” -
PNAS 2021 118, 31, e2104624118
Henrik Pinholt, Søren S.-R. Bohr, Josephine F Iversen, Wouter Boomsma, Nikos S Hatzakis*
“Single particle Diffusional fingerprinting: A machine learning framework for quantitative analysis of heterogeneous diffusion” -
Nature Commun 2019 10, 5655
Rasmus Peter Thomsen, Mette Galsgaard Malle, Anders Hauge Okhol, Swati Krishnan, Søren Schmidt-Rasmussen Bohr, Rasmus Schøler Sørensen, Friedrich Simmel, Nikos S. Hatzakis*, Jørgen Kjems*
“A large size-selective DNA nanopore with sensing applications” -
CELL 2018 175, 1856–1871 (outcover)
Stefano Stella, Pablo Mesa, Johannes Thomsen, Bijoya Paul, Pablo Alcon, Simon B. Jensen, Bhargav Saligram, Matias E. Moses, Nikos S. Hatzakis* and Guillermo Montoya*
“Conformational Activation of Cas12a Catalysis and Endonuclease Resetting”
Knud J. Jensen lab (KJJ lab)
Jensen is an internationally recognized chemist with leading expertise on solid-phase synthesis and medicinal chemistry. His expertise in bio conjugation chemistry allowed his group to develop self-assembling peptide-oligonucleotide conjugates, new methods for solid-phase synthesis and protein conjugation, as well new methods for conjugating carbohydrates and lipids. He has worked closely with Hatzakis on chemical design and generation of biomolecules for single-molecule studies.
Read more about the on the UCPH website.
Selected papers:
-
J Am Chem Soc 2023 145, 16771-16777
Vanessa Rück, Narendra Kumar Mishra, Kasper K. Sørensen, Mikkel B. Liisberg, Ane Beth Sloth, Cecilia Cerretani, Christian B. Mollerup, Andreas Kjaer, Chenguang Lou, Knud J. Jensen, and Tom Vosch*
“Bioconjugation of a DNA-Stabilized Silver Nanocluster to Peptides and Human Insulin by copper-free click chemistry” -
Bioconjugate Chem 2023 34(3), 518-528
Shunliang Wu, Mads Østergaard, Freja Fredholt, Niels Johan Christensen, Kasper K. Sørensen, Narendra K. Mishra, Hanne M. Nielsen, and Knud J. Jensen*
“Ca2+-Responsive Glyco-insulin” -
Nature Commun 2022 13, 76
Shankar Pandey, Shankar Mandal, Mathias Danielsen, Asha Brown, Changpeng Hu, Niels J. Christensen, Alina Kulakova, Shixi Song, Tom Brown, Knud J. Jensen, Jesper Wengel, Chenguang Lou*, and Hanbin Mao*
“Chirality transmission in macromolecular domains” -
Chem Bio Chem 2022 23(24), 31, e202200359
Knud J. Jensen*, Mikkel B. Thygesen, Kasper K. Sørensen, Dr. Shunliang Wu, Tuule Treiberg, Sanne Schoffelen
“Selective Acylation of Proteins at Gly and Lys in His Tags” -
Nature Commun 2016 7, 12294
Chenguang Lou, Manuel C. Martos-Maldonado, Charlotte S. Madsen, Rasmus P. Thomsen, Søren R. Midtgaard, Niels J. Christensen, Jørgen Kjems, Peter W. Thulstrup, Jesper Wengel* and Knud J. Jensen*
“Peptide-oligonucleotide conjugates as nanoscale building blocks for assembly of an artificial three-helix protein mimic”
Kirchhausen lab
Kirchhausen has pioneered our understanding of receptor-mediated endocytosis. He is a world leader in studying traffic of membrane proteins and cargo in cells, organelle biogenesis, and mechanisms of virus-host cell interactions, by use of frontier imaging methods -- most recently, lattice light sheet microscopy, recognized as a revolution in fluorescence imaging and 3D focused ion beam scanning microscopy combined with AI-based image processing. Relevant on-going work concerns delivery of lipid-encapsulated siRNA and naked antisense oligonucleotides (ASOs) into cells.
Read more about the Kirchhausen lab here.
Selected papers:
-
BioRxiv 2024 2023.12.30.573738 (preprint)
Anwesha Sanyal, Gustavo Scanavachi, Elliott Somerville, Anand Saminathan, Athul Nair, Athanasios Oikonomou, Nikos S. Hatzakis, Tom Kirchhausen*
“Constitutive Endolysosomal Perforation in Neurons allows Induction of α-Synuclein Aggregation by Internalized Pre-Formed Fibrils” -
J Cell Biol 2023 222(2), e202208005
Benjamin Gallusser, Giorgio Maltese, Giuseppe Di Caprio, Tegy John Vadakkan, Anwesha Sanyal, Elliott Somerville, Mihir Sahasrabudhe, Justin O’Connor, Martin Weigert, Tom Kirchhausen*
“Deep neural network automated segmentation of cellular structures in volume electron microscopy” -
BioRxiv 2023 2023.11.16.567393 (preprint)
Jacob Kæstel-Hansen, Marilina de Sautu, Anand Saminathan, Gustavo Scanavachi, Ricardo F. Bango Da Cunha Correia, Annette Juma Nielsen, Sara Vogt Bleshøy, Wouter Boomsma, Tom Kirchhausen*, Nikos S. Hatzakis*
“Deep learning assisted single particle tracking for automated correlation between diffusion and function” -
Developmental Cell 2021 56(12), 1786-1803.e9
Yi-Ying Chou, Srigokul Upadhyayula*, Justin Houser, Kangmin He, Wesley Skillern, Gustavo Scanavachi, Song Dang, Anwesha Sanyal, Kazuka G.Ohashi, Giuseppe di Caprio, Alex J. B. Kreutzberger, Tegy John Vadakkan, & Tom Kirchhausen*
“Inherited nuclear pore substructures template post-mitotic pore assembly” -
Nature 2017 552(7685), 410-414
Kangmin He*, Robert Marsland III, Srigokul Upadhyayula, Eli Song, Song Dang, Benjamin R. Capraro, Weiming Wang, Wesley Skillern, Raphael Gaudin, Minghe Ma & Tom Kirchhausen*
“Dynamics of phosphoinositide conversion in clathrin-mediated endocytic traffic”
Maria Carmo-Fonseca Lab
Maria Carmo-Fonseca is Professor at the University of Lisbon Medical School. She is a founder of the Institute of Molecular Medicine (iMM), a biomedical research institute affiliated with the University of Lisbon Medical School, where she currently serves as President. She was visiting Professor at Harvard Medical School (2011 to 2013). She is a member of the European Molecular Biology Organization, the Portuguese Academy of Sciences, the Portuguese Academy of Medicine, and Academia Europaea, and she served as President of the RNA Society (2021-2022). She has been scientific editor for the Journal of Cell Science and the RNA journal.
The Carmo-Fonseca lab combines microscopy techniques and genome-wide methodologies to study RNA splicing in health and disease.
Read more about the Maria Carmo-Fonseca Lab here.
Selected papers:
-
Nature Cell Biology 2023 25(4), 516-517
Maria Carmo-Fonseca*
“A twist to splicing regulation in haematopoiesis” -
CELL 2023 186(1), 10-11
Maria Carmo-Fonseca*
“Sweet splicing.” -
Molecular Cell 2022 82(5), 1021-1034.e8
Luna Tammer, Ofir Hameiri, Ifat Keydar, Vanessa Rachel Roy, Asaf Ashkenazy-Titelman, Noélia Custódio, Itay Sason, Ronna Shayevitch, Victoria Rodríguez-Vaello, José Rino, Galit Lev Maor, Yodfat Leader, Doha Khair, Erez Lieberman Aiden, Ran Elkon, Manuel Irimia, Roded Sharan, Yaron Shav-Tal, Maria Carmo-Fonseca, & Gil Ast*
“Gene architecture directs splicing outcome in separate nuclear spatial regions” -
Molecular Cell 2022 82(3), 495-496
Rui Sousa-Luís, & Maria Carmo-Fonseca*
“Pseudouridylation: A new player in co-transcriptional splicing regulation” -
RNA 2022 28(2), 139-161
Pedro Prudêncio, Rosina Savisaar, Kenny Rebelo, Rui Gonçalo Martinho*, Maria Carmo-Fonseca*
“Transcription and splicing dynamics during early Drosophila development”
Bi-annual symposium
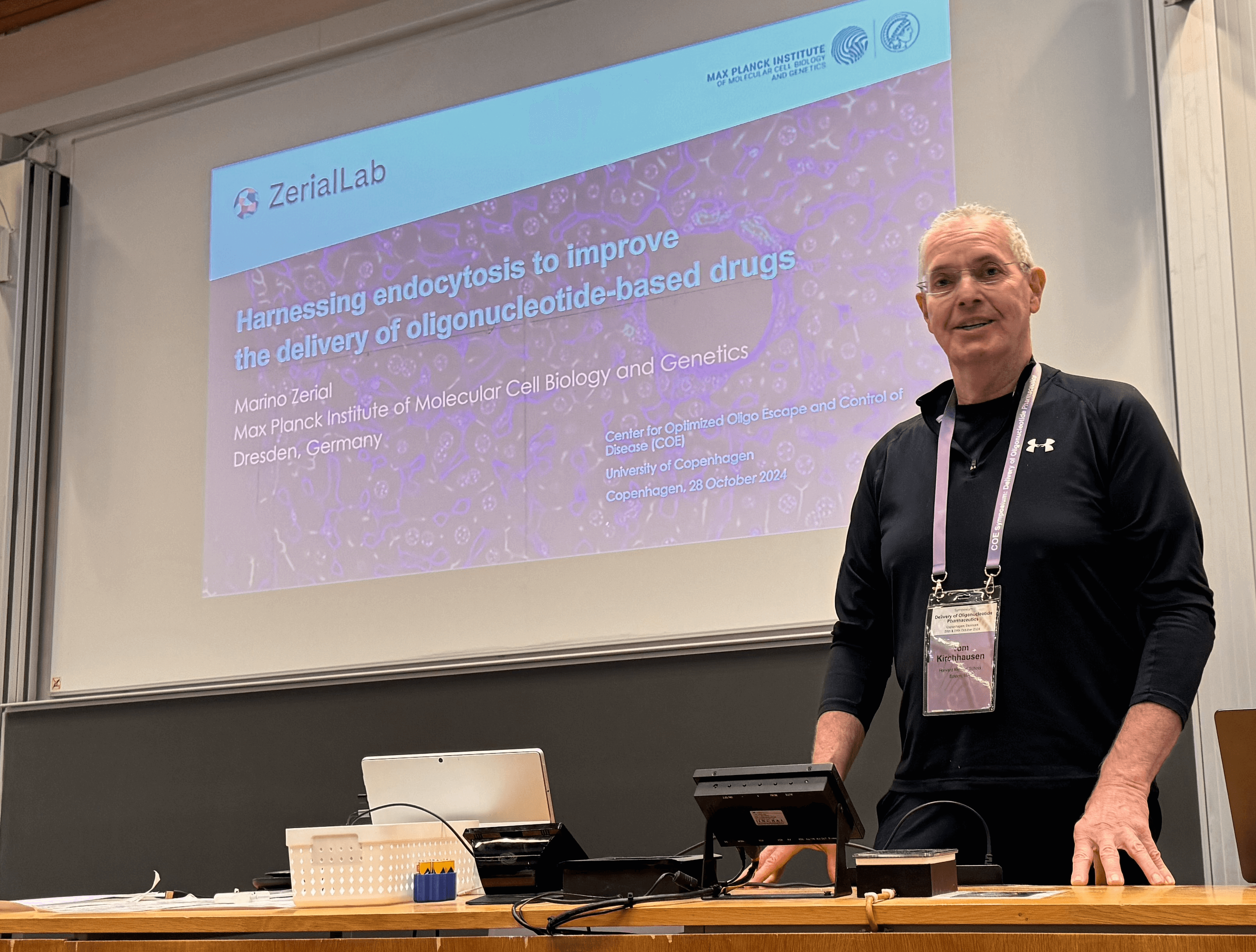
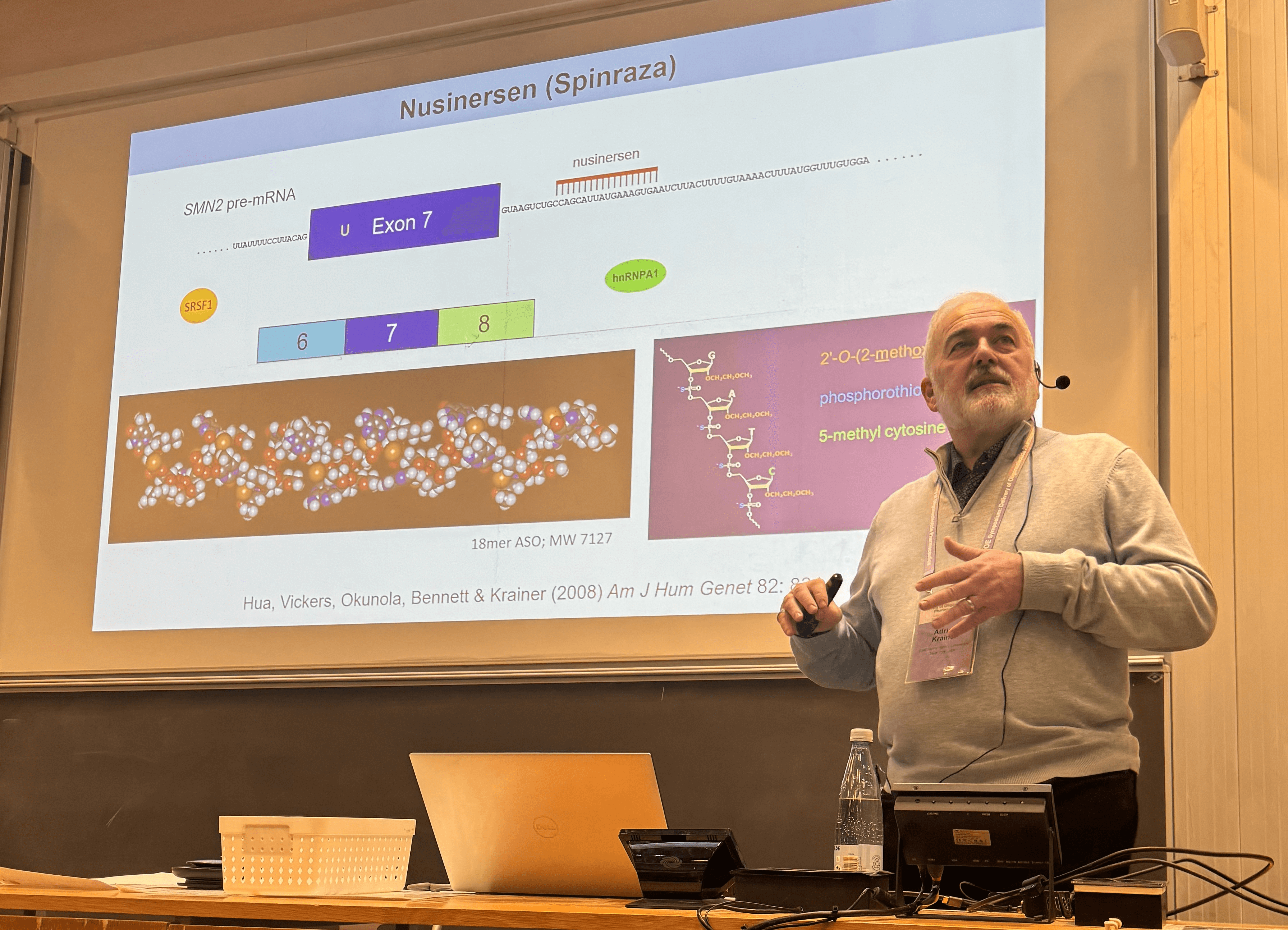
October 28 and 29, 2024, we held the COE symposium “Delivery of Oligonucleotide Pharmaceutics". We were delighted to welcome renowned speakers leading the respective field of oligonucleotide pharmaceutics. We were happy to see the high participation of both industry and academia.
Full program: Symposium on Delivery of oligonucleotide pharmaceutics.


Internal meetings
Every month, all COE members meet online to discuss projects, collaborations and science.
October 29 and 30, 2024, we also held our first internal on-site meeting after the COE symposium. We were happy to finally all meet in person and discuss collaborative projects within the center.

You can contact us with questions:
Nikos Hatzakis
Center leader
E-mail: Hatzakis@chem.ku.dk
Tania Darphorn
Center Coordinator
E-mail: tsd@chem.ku.dk
Mailing address Copenhagen:
Thorvaldsensvej 40, 1871 Frederiksberg C
University of Copenhagen
Copenhagen, Denmark
office: T554
Contact
Professor Nikos Hatzakis
Center leader
E-mail: Hatzakis@chem.ku.dk
Tania Darphorn
Center Coordinator
E-mail: tsd@chem.ku.dk
